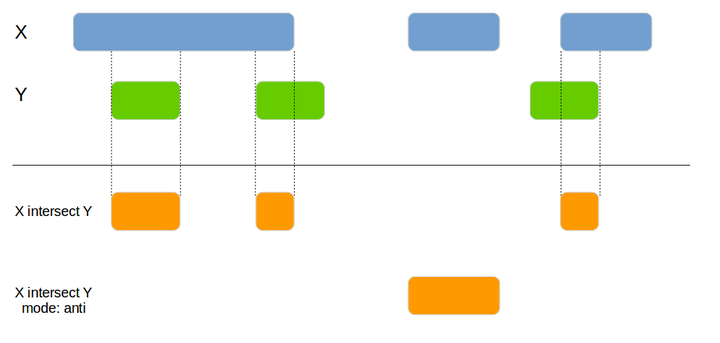
tidygenomics provides verbs for handling data frames that contain genomic locations.
x1 <- data.frame(id = 1:4,
chromosome = c("chr1", "chr1", "chr2", "chr2"),
start = c(100, 200, 300, 400),
end = c(150, 250, 350, 450))
x2 <- data.frame(id = 1:4,
chromosome = c("chr1", "chr2", "chr2", "chr1"),
start = c(140, 210, 400, 300),
end = c(160, 240, 415, 320))
genome_intersect(x1, x2, by=c("chromosome", "start", "end"), mode="both")
For more information see the tidygenomics CRAN page and the corresponding Github repository.