Constantin Ahlmann-Eltze
Postdoc
UCL London
Biography
I am a postdoc at the Cancer Institute at UCL in the group of James Reading. I did my PhD with Wolfgang Huber at EMBL Heidelberg. Before joining UCL, I did a bridging postdoc with Simon Anders at Heidelberg University.
Currently, I am working on novel ways to analyze the role of T cells in early carcinogenesis.
Previously, I have worked on:
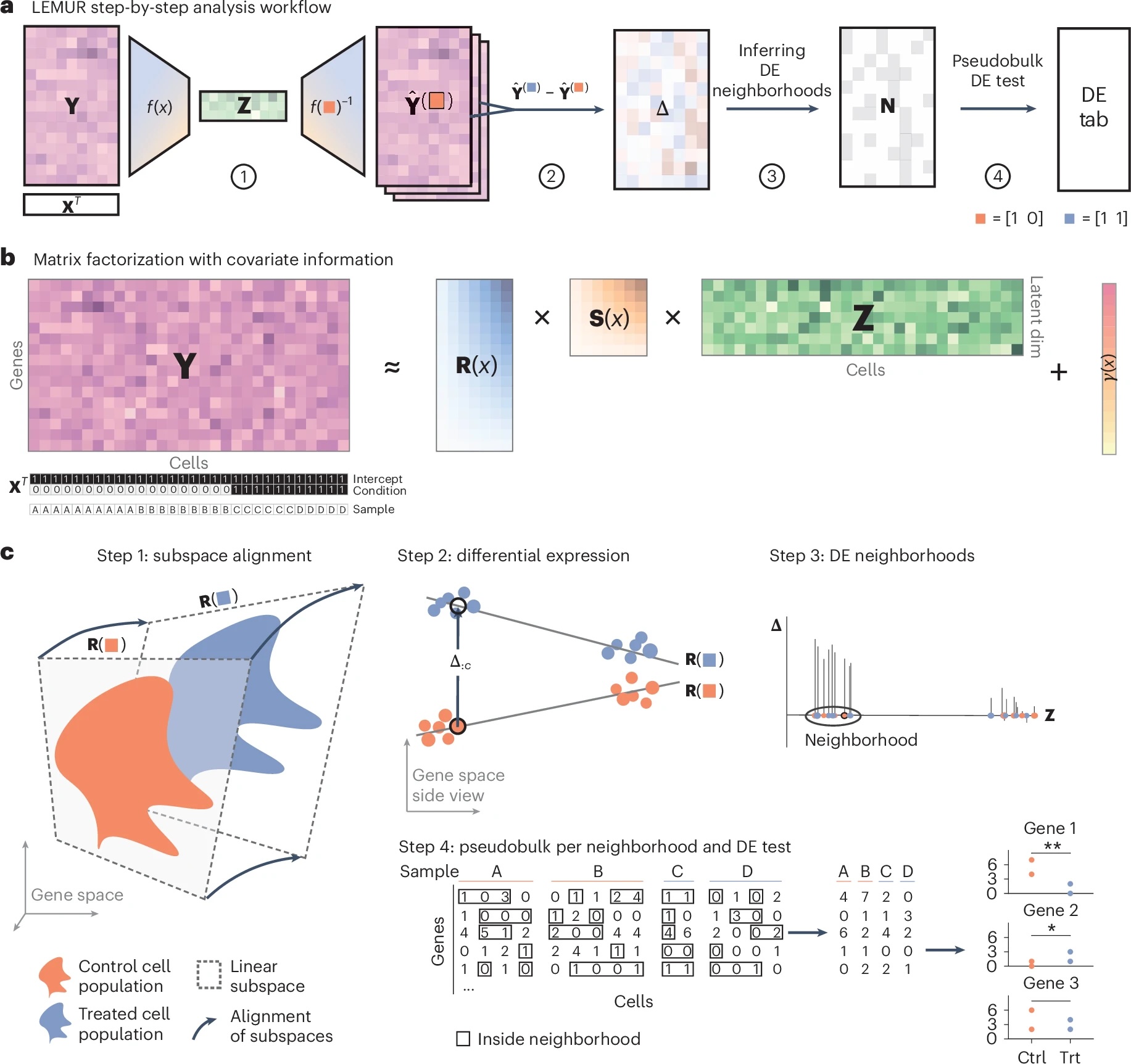
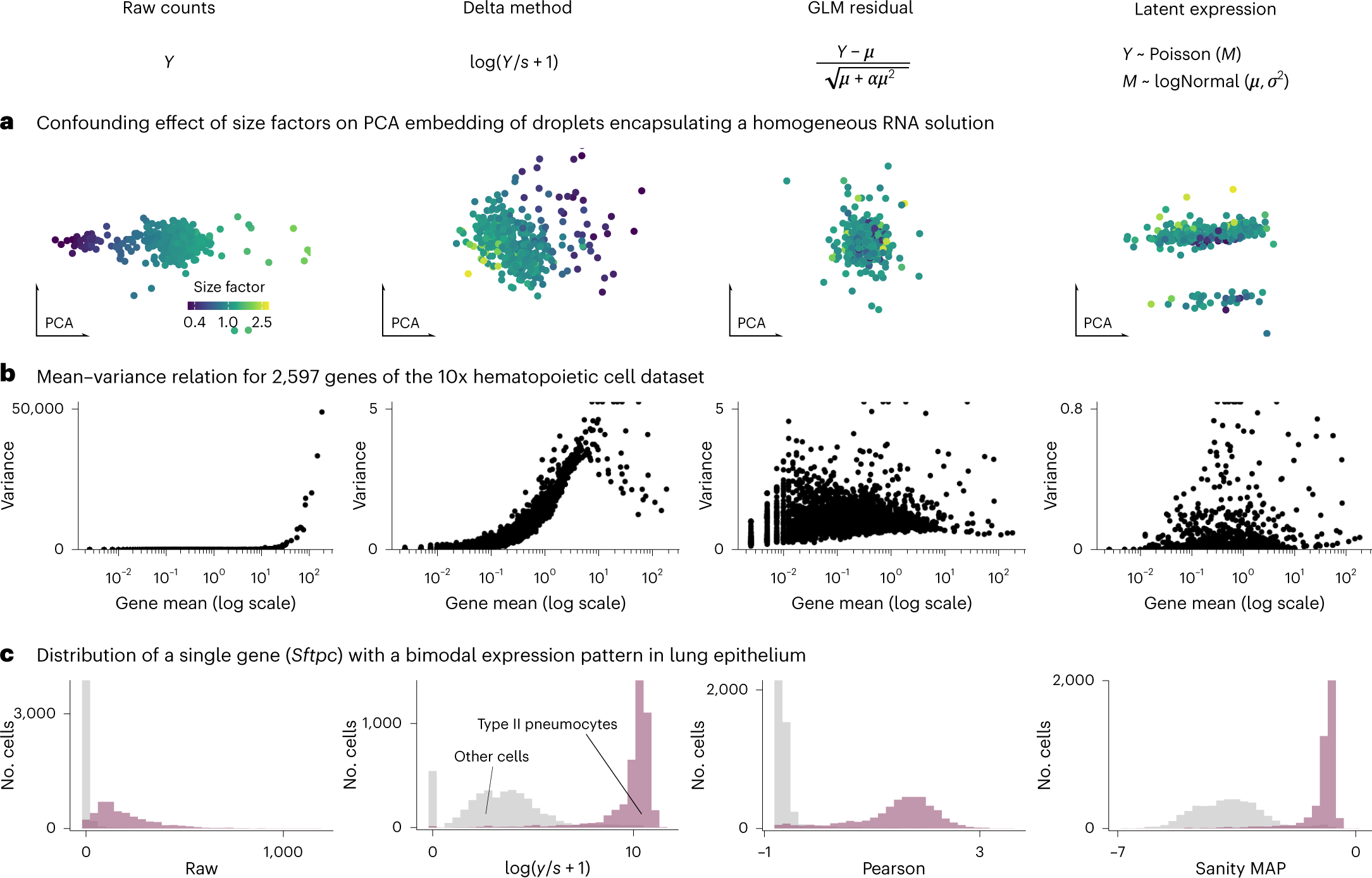
- Statistical methods to simplify working with single-cell data,
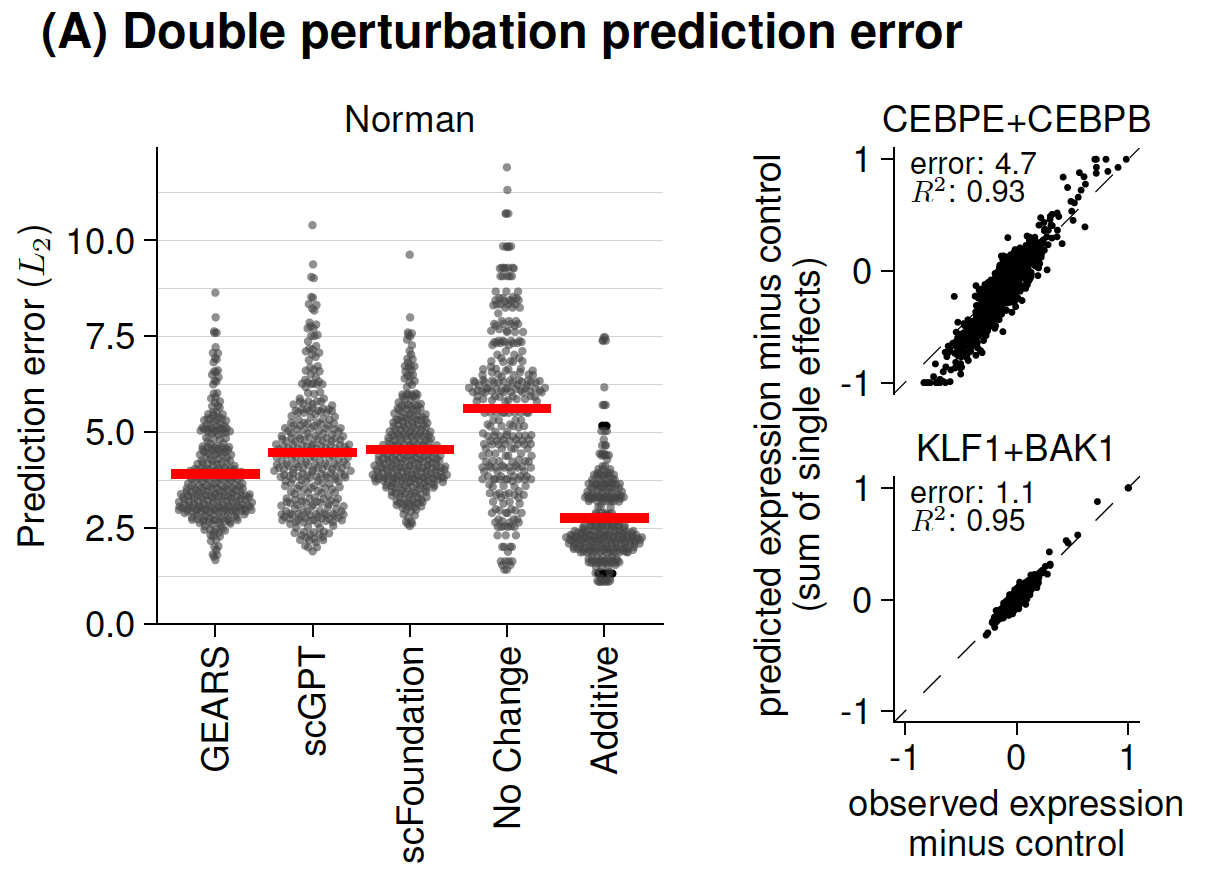
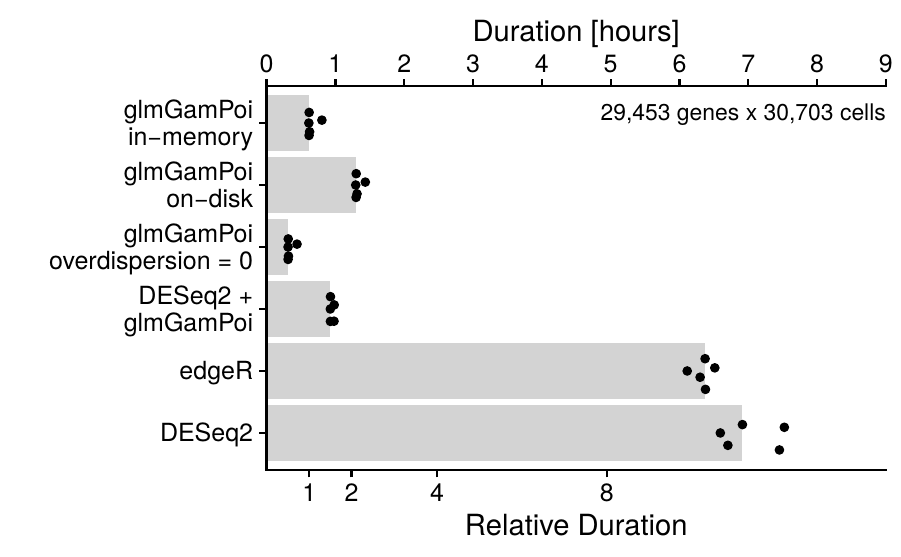
- Benchmarks of single-cell methods,
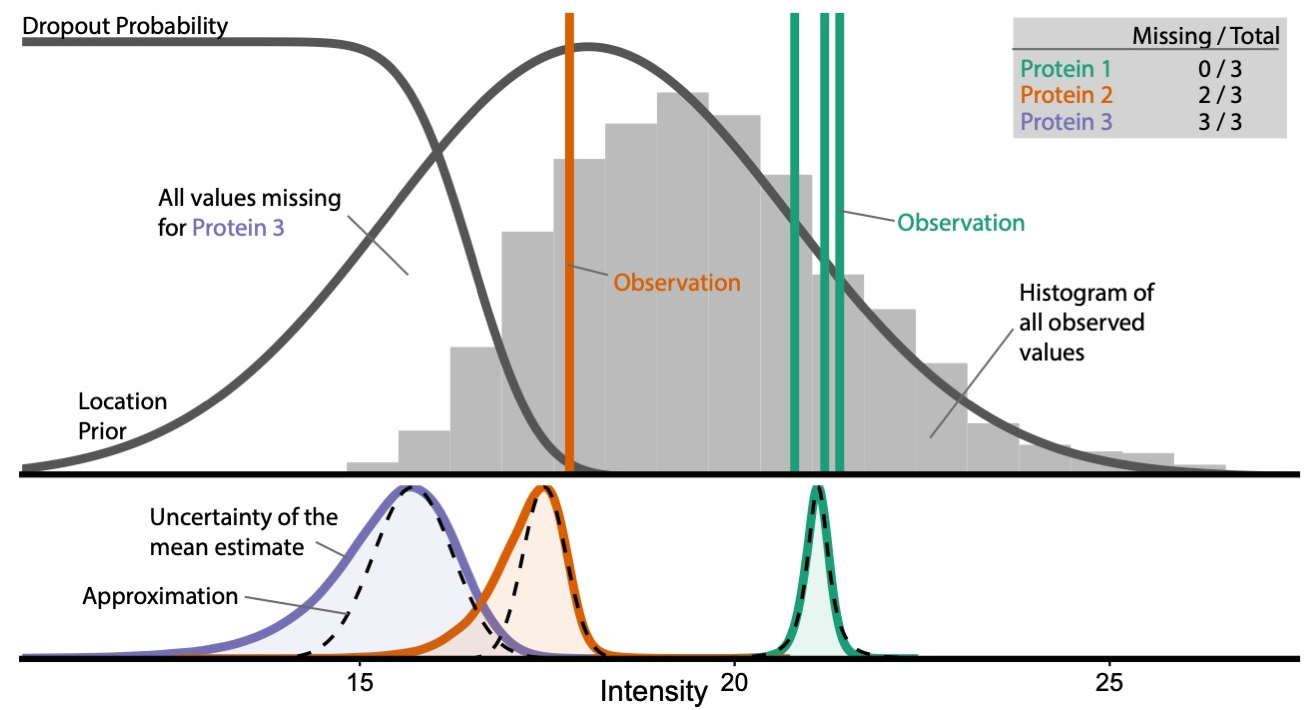
- Differential abundance analysis of mass spectrometry data,
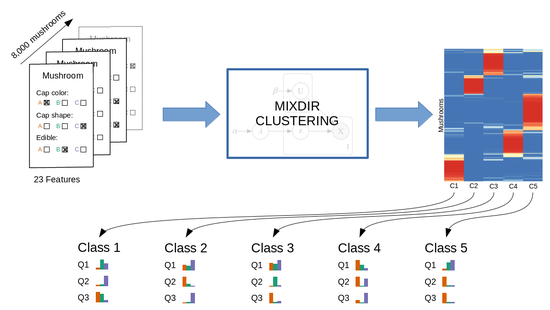
- Clustering of high-dimensional categorical data.
For a full record of my publications, see Google Scholar.
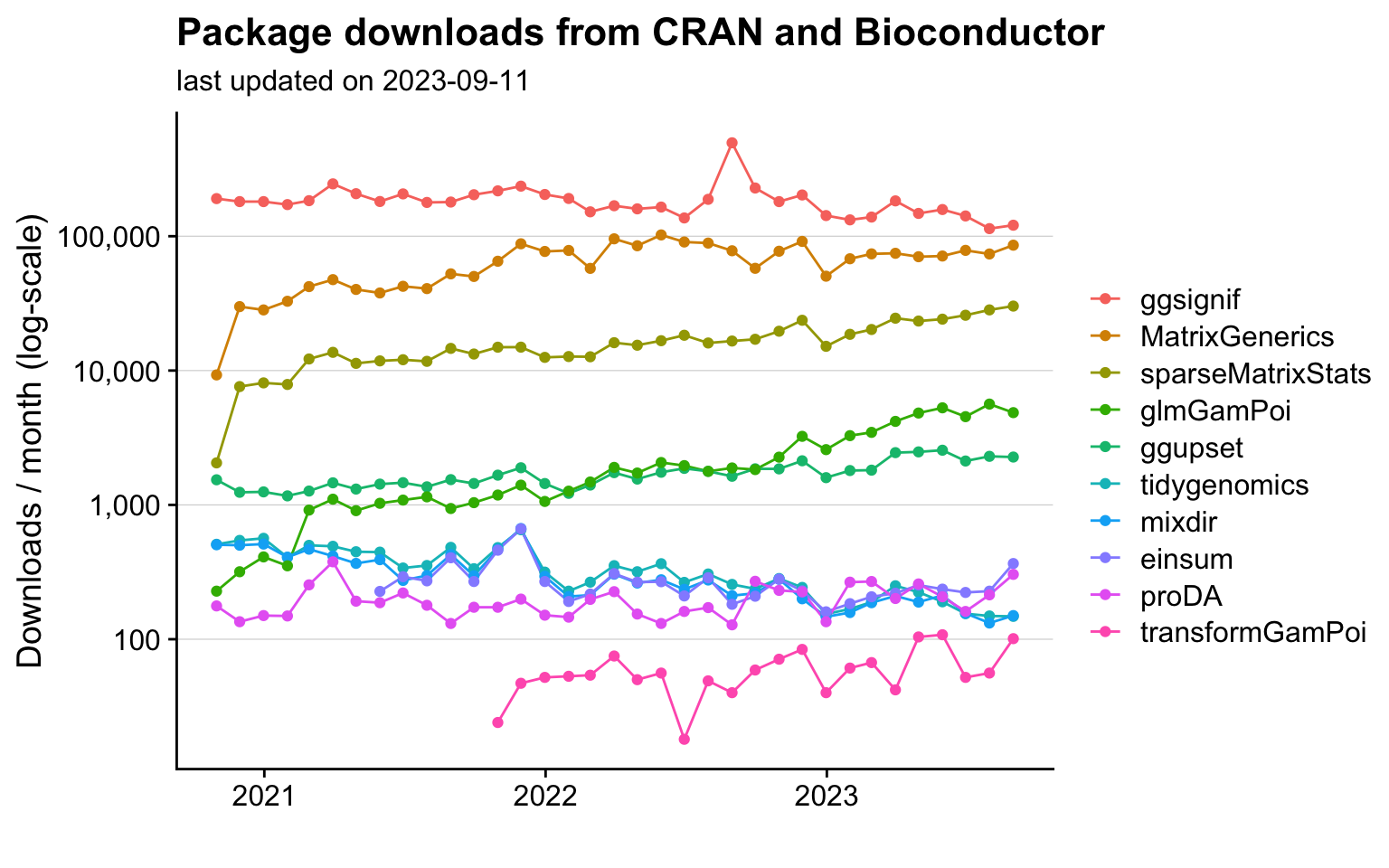
In addition, I develop statistical methods and tools for analyzing cutting-edge biological data. I maintain ten R packages, which are published on CRAN and Bioconductor and total more than 100,000 downloads per month. In 2023, I received the Bioconductor Community Award in recognition of my contributions to the project.
If you are working in a company and would like help with some analysis or one of my packages, don’t hesitate to get in touch, as I am happy to consult on projects.