proDA: Probabilistic Dropout Analysis for Identifying Differentially Abundant Proteins in Label-Free Mass Spectrometry
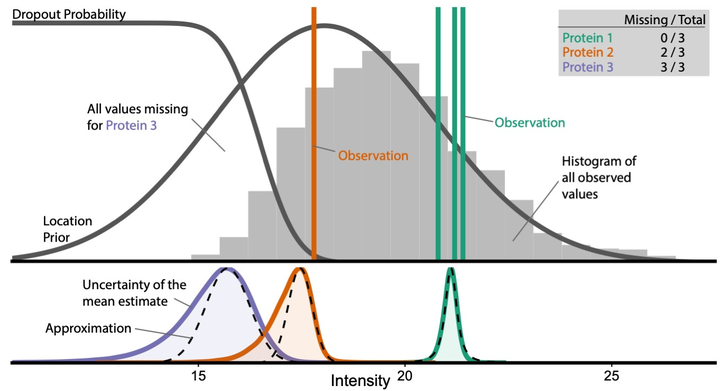 proDA Intuition
proDA Intuition
Abstract
Protein mass spectrometry with label-free quantification (LFQ) is widely used for quantitative proteomics studies. Nevertheless, well-principled statistical inference procedures are still lacking, and most practitioners adopt methods from transcriptomics. These, however, cannot properly treat the principal complication of label-free proteomics, namely many non-randomly missing values. We present proDA, a method to perform statistical tests for differential abundance of proteins. It models missing values in an intensity-dependent probabilistic manner. proDA is based on linear models and thus suitable for complex experimental designs, and boosts statistical power for small sample sizes by using variance moderation. We show that the currently widely used methods based on ad hoc imputation schemes can report excessive false positives, and that proDA not only overcomes this serious issue but also offers high sensitivity. Thus, proDA fills a crucial gap in the toolbox of quantitative proteomics. Availability: The proDA method is implemented as an open-source R package, available on https://github.com/const-ae/proDA